
|
AltaiR |
A C toolkit for alignment-free and spatial-temporal analysis of multi-FASTA data
|
University of Aveiro |
|
AlcoR |
Alignment-free simulation, mapping, and visualization of low-complexity regions in FASTA data.
|
University of Aveiro |

|
CoV2ID |
CoV2ID provides a complete, quality checked and regularly updated list of oligonucleotides for SARS-CoV-2. The database evaluates available therapeutic and detection protocols according to the virus genetic diversity.
|
CIIMAR |

|
WebSpecmine |
WebSpecmine is a web-based application designed to perform the analysis of metabolomics data based on spectroscopic and chromatographic techniques (NMR, Infrared, UV-visible, and Raman, and LC/GC-MS)) and compound concentrations.
|
University of Minho |

|
TriFusion |
Streamlining phylogenomic data gathering, processing and visualisation
|
|

|
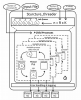
Structure_threader |
A wrapper program to parallelize and automate runs of "Structure", "fastStructure" and "MavericK".
|
|

|
SEDA |
SEDA (SEquence DAtaset builder) is an open source application for processing FASTA files containing DNA and protein sequences.
|
i3S - Institute for Research and Innovation in Health |
| |
PhagePromoter |
PhagePromoter is a python tool that predicts promoter sequences in phage genomes, using machine learning models.
|
University of Minho |

|
Smash++ |
|
University of Aveiro |

|
GTO |
GTO is a toolkit to unify pipelines in genomic and proteomic research.
|
University of Aveiro |

|
EAGLE |
EAGLE is an alignment-free method and associated program to compute relative absent words (RAW) in genomic sequences using a reference sequence.
|
University of Aveiro |

|
BLAST DataBase Manager |
BDBM (BLAST DataBase Manager) provides a graphical user interface that allows you to create high quality sequence datasets using the tools included.
|
i3S - Institute for Research and Innovation in Health |

|
ADOPS |
Bioinformatics tool that automatizes the detection of positively selected sites given a set of unaligned nucleotide sequence data
|
i3S - Institute for Research and Innovation in Health |

|
4Pipe4 |
A pipeline with emphasis on SNP detection. It will detect SNPs in NGS data where no reference sequence nor strain information is available.
|
Faculty of Sciences of the University of Lisbon |
|
NCBI Mass Sequence Downloader |
This program will download sequences en masse from several NCBI databases (at the user's choice).
|
Faculty of Sciences of the University of Lisbon |

|
MitoBreak: The mitochondrial DNA breakpoints database |
A comprehensive on-line resource with curated datasets of mitochondrial DNA (mtDNA) rearrangements. MitoBreak provides a complete, quality checked and regularly updated list of breakpoints from: circular deleted mtDNAs (deletions) circular partially-duplicated mtDNAs (duplications) linear mtDNAs
|
University of Porto |

|
COEUS |
Streamlined back-end framework for rapid semantic web application development.
|
University of Aveiro |

|
PHYLOViZ |
|
iMM - Instituto de Medicina Molecular João Lobo Antunes |

|
D-cellerate |
|
BioData.pt |

|
Bioinformatics Docker Images Project |
Dockerfiles for the images maintained by the Phenotypic Evolution Group at the i3S.
|
i3S - Institute for Research and Innovation in Health |

|
BioData.pt Data Stewardship Wizard |
Create, plan, collaborate, and bring your data management plans to life with a tool trusted by thousands of people worldwide — from data management pioneers, to international research institutes.
|
BioData.pt |
|
Biome-Shiny |
|
BioData.pt |








